Summary
Over the last 20-plus years, US Court of Appeals for the Federal Circuit cases concerning written description and enablement have become a hot-button issue in the chemical and life sciences practices. The year 2021 was no different, with Amgen v. Sanofi1 (enablement) decided in February and Juno v. Kite2 (written description) decided in August.3 Both Amgen and Juno involved genus claims with functional language, and both cases seemingly exacerbated the uphill battle for patent applicants and patentees to obtain and defend such claims. Indeed, the Federal Circuit’s recent stance on § 112 for chemical and life science genus claims has caused some to feel that the “sky is falling.”4 In light of the current § 112 landscape at the Federal Circuit, we examined the USPTO Patent Trial and Appeal Board’s (PTAB) recent views on written description and enablement law in the chemical and life sciences. For this work, we reviewed PTAB decisions from Technology Center 1600 (Biotechnology & Organic Chemistry), issued between January 2020 and November 2021. Our review included decisions from ex parte appeals and America Invents Act (AIA) post-grant proceedings (including decisions on institution and final written decisions).
We found that the sky is not falling, at least not at the PTAB. For example, we identified several recent ex parte appeals in which patent applicants successfully obtained broad genus claims after the PTAB’s reversal of Examiners’ § 112 written description and/or enablement rejections. The PTAB also – at least in some cases – considered evidence of routine screening to favor enablement of genus claims. While none of the PTAB cases highlighted below is currently designated precedential or informative, they nonetheless indicate that the PTAB’s application of § 112 offers patent owners in the chemical and life sciences a glimmer of hope. Below we highlight cases from our review, placing the PTAB decisions into three primary categories: (i) written description cases applying the representative species / common structural features rubric; (ii) written description cases applying the Capon factors; and (iii) enablement cases applying the routine screening rubric.
Written description: cases applying representative number of species / common structural features rubric.
In AbbVie Deutschland v. Janssen Biotech, 759 F.3d 1285 (Fed. Cir. 2014)—an antibody case involving genus claims with functional language—the Federal Circuit applied the representative number of species / common structural features rubric:
[A] sufficient description of a genus . . . requires the disclosure of
either a representative number of species falling within the scope of
the genus or structural features common to the members of the
genus so that one of skill in the art can ‘visualize or recognize’ the
members of the genus.
AbbVie, at 1299.
In August 2021, the Federal Circuit applied this rubric in Juno Therapeutics, Inc. v. Kite Pharma, Inc., 10 F.4th 1330 (Fed. Cir. 2021). The asserted claims in Juno were drawn to a nucleic acid polymer encoding a chimeric antigen T-cell receptor (CAR T-cell) comprising three segments: an intracellular signaling segment, a co-stimulatory segment comprising a specific amino acid sequence, and a binding segment (scFv). Id., at 1334. Juno’s specification disclosed two exemplary scFvs that bind to specific targets (CD19 and PSMA). Id., at 1333. Juno argued that other scFvs were known in the art, and that scFvs were interchangeable components with a shared, common structure. Id., at 1336.
The Federal Circuit acknowledged that scFv sequences were known in the art, and that scFvs share a common structure (seemingly satisfying the common structural features rubric), but nevertheless concluded that Juno’s specification lacked written description. The court stated:
[T]he written description of the ’190 patent discloses only two scFv
examples and provides no details regarding the characteristics,
sequences, or structures that would allow a person of ordinary skill
in the art to determine which scFvs will bind to which target. That
scFvs in general were well-known or have the same general
structure does not cure that deficiency.
Juno at 1339-1340 (emphasis added). In other words, as the court stated: “For the claimed functional scFv genus, the ’190 patent does not disclose representative species or common structural features to allow a person of ordinary skill in the art to distinguish between scFvs that achieve the claimed function and those that do not.” Id. at 1342.
How is the PTAB applying representative species / common structural features?
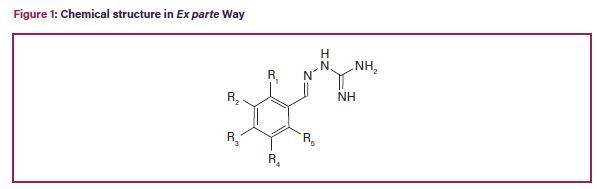
While the outlook for some biotech patent owners at the Federal Circuit may seem bleak at the moment, patent applicants continue to successfully obtain genus claims. For example, in Ex parte Way, No. 2019-006053 (PTAB, July 9, 2020), the claims were drawn to a “method of treating a demyelinating disorder” comprising administering a compound “of Formula I” (see Figure 1), “wherein R1, R2, R3, R4, and R5 are independently hydrogen, deuterium, halogen, haloalkyl, alkyl, alkoxy, hydroxyl, aryl, or aryloxy.” Way, at 2. Way’s specification disclosed working examples using a single compound (guanabenz) within the scope of the claimed genus of compounds. The Examiner rejected the claims for lack of written description and enablement. Id., at 3. On appeal, the PTAB reversed the Examiner’s written description rejection,5 stating that Way’s specification provided structural limitations that correlated with the claimed function: “Claim 2 recites a reasonably small genus of compounds with specific and complete structural limitations that are correlated with the function of treating a demyelinating disorder.” Way, at 9 (emphasis added).
Applicants in the biologics space have also successfully overturned § 112 rejections at the PTAB. In Ex parte Keler, No. 2019-006094 (PTAB, June 10, 2020), the applicant claimed a “method for inducing or enhancing an immune response” using a monoclonal antibody which binds to human CD27, “wherein the antibody comprises heavy and light chain variable region sequences having at least 95% identity to SEQ ID NOs: 37 and 43, respectively.” Id., at 2. Keler’s specification disclosed three exemplary antibodies with the claimed 95% sequence identity, and further disclosed data pertaining to binding, blocking, competition, and complement-mediated cytotoxicity. On appeal, the PTAB reversed the Examiner’s written description rejection, holding that the three example antibodies exhibit a common structure that correlates with the claimed function, thus satisfying written description:
The Specification further demonstrates that
these species exhibit both the structure recited
in claim 10 (i.e., comprises heavy and light chain
variable region sequences having at least 95%
identity to SEQ ID NOs: 37 and 43) as well as the
recited function (i.e., they bind to human CD27 and
induce/enhance an immune response) . . . . Thus,
Appellant’s description correlates the structure of
these species to the claimed function.
Keler, at 11 (emphasis added).
Ex parte Campbell, No. 2021-000865 (PTAB, July 20, 2021), is similar to Way. Campbell’s claims recited “a method of treating an autoimmune disease or condition, a systemic inflammatory disease or condition, or transplant rejection” comprising administering an anti-OX40L antibody having 90% sequence identity in the antibody’s heavy and light chain variable regions to specific sequences disclosed in the application. Id. at 2-3. Campbell’s specification disclosed two example antibodies, both meeting the claimed sequence identity limitations, and further disclosed experimental data such as ligand/receptor neutralization and IL-2 secretion assays. Id., at 9, 15.
On appeal, the PTAB reversed the Examiner’s §112 rejection, holding that the specification disclosed a representative number of species and that the claimed genus shared common structural features:
“[T]he claimed antibodies are not exclusively
claimed functionally, but are also claimed structurally,
i.e., by requiring that the light and heavy chains
have a structure corresponding to 90–95% of the
recited SEQ ID NOS. Furthermore, the number of
possible substitutions is relatively small: the variable
regions of the antibodies comprise approximately
100 amino acid sequences, and so the total number
of potential substitutions is no more than 10–12 . . .
Additionally, the Specification provides at least two
embodiments antibodies: 2D10 and 10A07, that are
fully functional in their antigen-binding capabilities.
Campbell, at 15 (emphasis added).
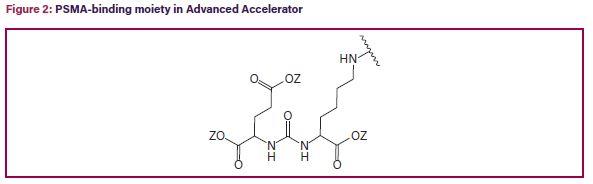
Recent PTAB decisions from AIA proceedings appeared to include more mixed outcomes when it comes to §112. One reason may be that AIA trials, unlike ex parte appeals, are inter partes proceedings where a motivated adversary presses the patentability issues. In the PTAB’s decision on institution in Advanced Accelerator Applications v. Molecular Insight Pharms, PGR2021-00048, Paper 7 (PTAB, July 29, 2021), the challenged claims recited a method of treating a patient with prostate cancer comprising administering a therapeutically effective amount of a glutamate-urea-lysine PSMA-binding moiety comprising the structure shown in Figure 2, wherein each Z, independently, is H or C1-C4 alkyl. Id., at 4. The specification disclosed in vitro binding data (IC50 values) for about two dozen compounds, and provided biological data for a single compound. Id., at 18, 24.
At the institution stage, the PTAB instituted post-grant review because it determined it was more likely than not that the challenged claims lacked written description:
[T]he ’461 patent does not disclose a sufficiently
representative number of species because the
patent only provides PSMA-binding data for a
handful of compounds, many of which show poor
binding, and only provides further biological in
vivo testing for one compound, MIP-1072. Given
the apparent breadth of the challenged claims,
this limited number of disclosed compounds and
limited data does not appear sufficient to provide
adequate written description support. We also
find that the record sufficiently shows that the
’461 patent does not disclose structural features
common to members of the genus.
Id. at 24. Although Advanced Accelerator is still in trial,6 the decision at institution illustrates that the PTAB is carefully scrutinizing the representative species/common structural features rubric. And we found several other PTAB AIA decisions applying rationales similar to Advanced Accelerator. See e.g., Syngenta Crop Protection AG v. FMC Corporation, PGR2020-00028, Paper 33 (PTAB, Aug. 31, 2021) (finding claims covering “more than a billion different compounds” unpatentable for lack of written description, when the specification failed to “divine a relationship between structure and activity, given that the test data is clustered around a narrow range of structures.”); Allgenesis Biotherapeutics v. Cloudbreak Therapeutics, IPR2020-01438, Paper 7 (PTAB, Feb. 18, 2021) (“disclosure of a single species [of multikinase inhibitor] cannot be extrapolated to the genus.”).
In contrast to Advanced Accelerator, the PTAB determined in SweeGen, Inc. v. PureCircle Sdn Bhd, PGR2020-00070, Paper 14 (Jan. 19, 2021), that the petitioner failed to show it was more likely than not to prevail on its §112 written description arguments. In SweeGen, the claims recited a method of adding a glucose unit to a steviol glycoside comprising contacting the steviol glycoside with an enzyme comprising UDP-glucosyltransferase. The PTAB rejected the petitioner’s assertion that the claimed scope was overbroad in view of the specification: “We agree with Patent Owner that a person of ordinary skill in the art could envision the steviol glycosides and UDP-glucosyltransferases encompassed by the claims because they have common structural features.” SweeGen, at 32 (emphasis added). Thus, in contrast to Advanced Accelerator, the PTAB panel in SweeGen noted that members of the genus shared common structural features.
This snapshot of PTAB decisions in TC1600 indicates that the PTAB seems to apply the representative number of species / common structural features rubric in a relatively balanced manner that, at least in some cases, favors the patent applicant or patentee. Time will tell if the PTAB begins applying a stricter standard in light of Juno.
Written description: cases applying the Capon factors
We also reviewed PTAB decisions to assess how the PTAB applied Capon v. Eshhar, 418 F.3d 1349 (Fed. Cir. 2005). Capon established the following well-known factors for assessing written description of a genus claim in the biological arts:
[T]he determination of what is needed to support
generic claims to biological subject matter
depends on a variety of factors, such as the existing
knowledge in the particular field, the extent and
content of the prior art, the maturity of the science
or technology, the predictability of the aspect at
issue, and other considerations appropriate to the
subject matter.
Capon, at 1359. Since Capon was decided in 2005, patentees and patent applicants alike have often interpreted it as confirming that common knowledge in the art—such as known nucleotide sequences—need not be disclosed in the specification to support written description.
The Federal Circuit in Juno acknowledged this principle, stating that a patentee need not “in all circumstances” disclose nucleotide or amino acid sequences “when such sequences are already known in the prior art.” Juno, at 1337. However, the Court in Juno distinguished Capon:
Our Capon decision neither made the
determination Juno alleges nor determined
that the inventors there satisfied the written
description requirement. Instead, we vacated the
Board’s decision for imposing too high a standard
to satisfy the written description requirement,
and remanded for the Board to consider the
evidence and determine whether the specification
adequately supported the claims at issue . . . Capon
does not support Juno’s arguments regarding its
exceedingly broad functional claim limitations.
Id., at 1338 (emphasis added). With this new insight from Juno, we investigated how the PTAB is applying Capon.
How is the PTAB applying Capon?
Juno does not appear to have impacted the PTAB’s stance on Capon in TC1600—at least not yet. Indeed, we found PTAB decisions issued before and after Juno that applied the same interpretation of Capon. For example, in Ex parte Harriman, No. 2020-004459 (PTAB, Feb. 10, 2021), decided before Juno, the claims recited a transgenic chicken comprising human immunoglobulin genes. The specification did not disclose any specific chicken or human immunoglobulin sequences, and provided only prophetic examples. The Examiner rejected the claims for lacking written description, asserting that the specification did not disclose any “transgene comprising an exogenous ‘pre-arranged human light chain Ig gene’ for targeted integration.” Harriman at 8. In reversing the §112 rejection, the PTAB expressly relied on Capon, stating that “a pre-rearranged human Ig light chain variable region simply requires knowledge of human Ig light chain variable region sequences, which are replete in Genbank and other sources.” Id. at 11. The PTAB explained: “Consistent with Capon, the ordinary artisan may select any known described deposited sequences for joinder by PCR or other well-known methods and use in the invention.”
Id.; see also, Ex parte Roninson, No. 2019-006086 (PTAB, June 9, 2020) (applying Capon in reversing Examiner’s §112 rejection); Ex parte Terbrueggen, No. 2018-004820 (PTAB, April 1, 2020) (applying Capon in reversing Examiner’s §112 rejection; affirming on other grounds).
In Ex parte Oliver, No. 2021-000044 (PTAB, Oct. 8, 2021), decided after Juno, the claims recited a method for preparing a biomolecule analyte comprising hybridizing oligonucleotide probes to a single-stranded human DNA or human RNA template, performing a base extension reaction (e.g., PCR), terminating the reaction such that a single-strand region is adjacent to a hybridized probe, and reacting the product with a binding moiety. The Examiner argued that the specification did not provide any specific sequences for the template or oligonucleotide probes, and did not describe, e.g., how to direct a probe to polyA sequences that are not adjacent to one another. Id., at 9-10. The PTAB rejected the Examiner’s hypothetical, and, citing Capon, stated that a skilled artisan would have relied on the general knowledge in the art:
[W]e are not persuaded that Examiner’s postulated
hypothetical, which would have been recognized
as inoperable by those of ordinary skill in this
art, at the time of Appellant’s claimed invention,
supports a conclusion that Appellant’s claimed
invention lacks written descriptive support. See,
e.g., Capon, 418 F.3d at 1359 (‘It is not necessary
that every permutation within a generally operable
invention be effective in order for an inventor to
obtain a generic claim.’)
Oliver, at 10 (emphasis added).
Similarly, in Ex parte Landegren, No. 2021-001167 (PTAB, Nov. 17, 2021), decided after Juno, the claims recited a method of selecting a target region of interest (ROI) in a target nucleic acid, comprising a specific oligonucleotide probe capable of hybridizing with itself to form a stem loop structure, and a series of steps involving hybridizing a probe to the target nucleic acid, base extension reactions (e.g., PCR), ligations to circularize the extended probes, and further amplification. The Examiner rejected the claims under §112(a), asserting that the probes comprise multiple sequences that can bind to multiple undisclosed targets, that the ROI can range from 10 to 100,000 nucleotides in length, but the specification only discloses three example target sequences. Landegren, at 4-5. In reversing the §112 rejection, the PTAB cited Capon and leaned on the knowledge in the art:
In view of the state of the art and the knowledge
of those working in the areas such as nucleic
acid hybridization, ligation, and amplification, the
Examiner has not shown that the description
provided by the Specification of the claimed
method, and of the probes used in it, would fail to
show possession to those of ordinary skill in the art.
Landegren, at 9 (emphasis added).
While this sample size is small, it appears—at least for now—that Juno has not had much of an impact at the PTAB.
Enablement: cases applying the routine screening rubric
Enablement cases at the Federal Circuit are also a topic of debate lately. In Idenix Pharmaceuticals LLC v. Gilead Sciences Inc., 941 F.3d 1149 (Fed. Cir. 2019), the claims at issue recited a method of treating hepatitis C virus (HCV) infection with a particular type of nucleoside compound. Idenix argued that the key to its invention and the treatment of HCV infection is the use of “2’-methyl-up” nucleosides. Idenix, at, 1154. Gilead argued that Idenix’s claim was overbroad, and Idenix’s patent specification provided no guidance in determining which of the “billions and billions” of potential 2’-methyl-up nucleosides are effective in treating HCV. Id., at 1157. Idenix’s specification provided four examples. Id., at 1161. The Court held that, even though synthesis of the 2’-methyl-up compounds was routine, “[T]he immense breadth of screening required to determine which 2’-methyl-up nucleosides are effective against HCV can only be described as undue experimentation.” Id., at 1162. The Court affirmed the district court’s judgment that Idenix’s claims were invalid for lack of enablement. Id., at 1165.
In February 2021, the Federal Circuit decided Amgen Inc. v. Sanofi, Aventisub LLC, 987 F.3d 1080 (Fed. Cir. 2021). Amgen’s claims recited an antibody that binds at least one or at least two specific amino acid residues in the PCSK9 receptor and blocks binding of PCSK9 to its ligand, LDLR. The Federal Circuit noted that functional claim limitations “pose high hurdles in fulfilling the enablement requirement for claims with broad functional language” and “the use of broad functional claim limitations raises the bar for enablement.” Amgen, at 1087 (emphasis added). Similar to Idenix, the Federal Circuit in Amgen held that the skilled artisan must be able to make and screen every antibody within the genus for the claims to be enabled, explaining that “the scope of the claims encompasses millions of candidates claimed with respect to multiple specific functions, and that it would be necessary to first generate and then screen each candidate antibody to determine whether it meets the double-function claim limitations.” Amgen, at 1088 (emphasis added). The Court affirmed the district court’s judgment that Amgen’s claims were invalid for lacking enablement. Id.
How is the PTAB applying routine screening for enablement?
We found varying views from the PTAB on routine screening and enablement. For example, in Ex parte Way, supra, the PTAB assessed and weighed each of the Wands7 factors, noting that the specification disclosed routine screening assays that weighed in favor of enablement. Way, at 7. Similarly, in SweeGen, supra, the PTAB again assessed the Wands factors to find that the petitioner failed to establish it was more likely than not to prevail on its enablement challenge. There, the PTAB stated that, “although there is some unpredictability in the art, the testing appears to have been routine and information known in the art as well as homology modeling could have been used to make the field somewhat more predictable and reduce the amount of experimentation needed.” SweeGen, at 24 (emphasis added).
However, in Advanced Accelerator, supra, the PTAB leaned into Idenix and Amgen, stating that “the sheer number of candidate compositions that must be synthesized and then assayed weighs against enablement. This is especially true in light of the lack of guidance in the specification as to which of these compounds would have such therapeutic activity.” Advanced Accelerator, at 20. Likewise, in Syngenta Crop Protection, supra, the PTAB acknowledged that synthesis and screening of compounds was routine, but—relying on Idenix—concluded that the volume of synthesis and screening weighed against enablement: “[S]imilar to Idenix, despite the high level of skill in the art and routine nature of synthesis and screening techniques, the ‘immense breadth of screening required to determine which [compounds] are effective [herbicides] can only be described as undue experimentation.’” Syngenta, at 40 (quoting Idenix at 1162).
Finally, in Genome & Co. v. University of Chicago, PGR2019-00002, Paper 40 (PTAB, April 14, 2020), the PTAB found the claims unpatentable for lack of enablement due, in part, to the specification’s focus on antibodies. In Genome, the claims recited a method of treating cancer that involved administering “an immune checkpoint inhibitor,” but did not recite any specific type of checkpoint inhibitor. In finding the claims unpatentable for lacking enablement, the PTAB stated, “While the Specification defines [checkpoint inhibitors] broadly as including a protein or polypeptide that binds an immune checkpoint as well as an interfering nucleic acid molecule, the CPIs listed in the Specification are almost exclusively antibodies.” Genome, at 15. The PTAB explained, “the Specification gives no guidance as to how to select a CPI that, other than those recited in the Specification, is useful in the practice of the invention.” Id., at 28.
Some of these recent PTAB decisions may sound similar to Idenix or Amgen in that the PTAB considered the breadth of the claimed genus to outweigh the benefits of routine synthesis and screening for enablement. See e.g., Advanced Accelerator; Syngenta. Patent applicants and owners, however, need not give up on arguing that routine screening favors enablement, as some panels deciding cases arising from TC1600 still found that routine screening weighed in favor of enablement of a genus claim. See e.g., Way; SweeGen.
Conclusion
While recent decisions from the Federal Circuit have arguably increased § 112 scrutiny for patent owners in the chemical and life sciences, the sampling of PTAB cases in TC1600 presented here indicates that perhaps the sky is not falling; at least, not at the PTAB. At least not yet. Many patent applicants and patentees were able to successfully obtain (or defend) genus claims at the PTAB. It remains to be seen whether such claims can withstand scrutiny in litigation, but there is nevertheless value in genus claims even without asserting them in litigation. For example, genus claims can be valuable tools in licensing negotiations, attracting investors, asset sales/acquisitions, or simply serving as public notice to competitors.
1.Amgen Inc. v. Sanofi, Aventisub LLC, 987 F.3d 1080 (Fed. Cir. 2021), petition for cert. filed, _____ (U.S. Nov. 22, 2021) (No. 21-757).
2.Juno Ther., Inc. v. Kite Pharma, Inc., 10 F.4th 1330 (Fed. Cir. 2021).
3. The Federal Circuit also issued two decisions concerning written description in November 2021: Indivior UK v. Dr. Reddy’s Labs., 2020-2073 (Fed. Cir. Nov. 24, 2021) (addressing description of claimed ranges); and Biogen Int’l v. Mylan Pharms. Inc., 2020-1933 (Fed. Cir. Nov. 30, 2021 (addressing description of a “therapeutically effective amount”).
4. Amgen Inc. v. Sanofi, Aventisub LLC, 987 F.3d 1080 (Fed. Cir. 2021), rehearing denied, 850 F.App’x 794, 796 (Fed. Cir. 2021).
5. Discussed further below, the PTAB in Way also reversed the enablement rejection.
6. According to a Joint Stipulation from the parties, the Patent Owner’s Response is due January 21, 2022.
7.In re Wands, 858 F.2d 731 (Fed. Cir. 1988).
This article appeared in the 2021 PTAB Year in Review: Analysis & Trends report. To view our graphs on Data and Trends, please click here.
Related Industries

Receive insights from the most respected practitioners of IP law, straight to your inbox.
Subscribe for Updates